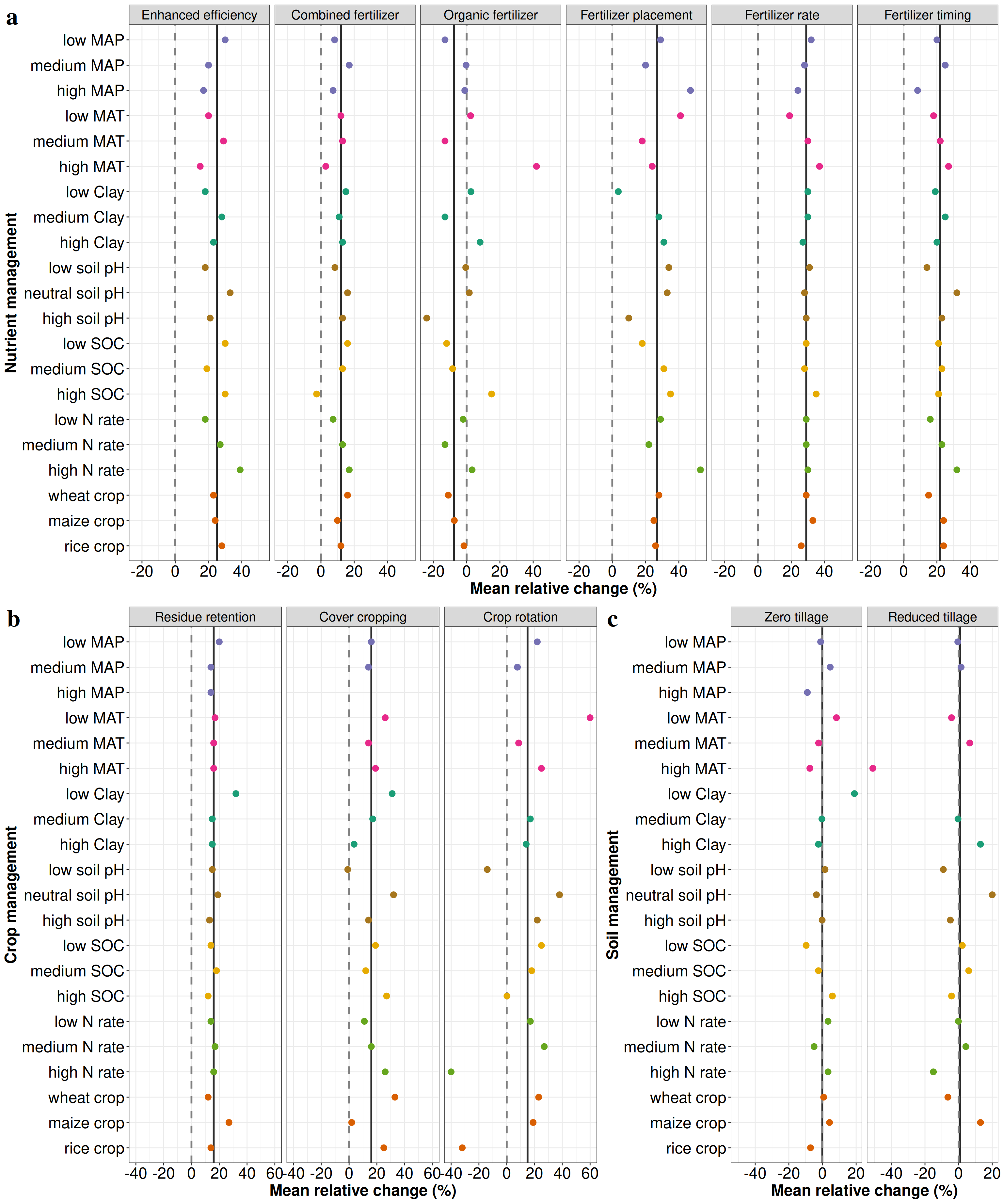
# Figure S4
library(dplyr)
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
Loading required package: ggplot2
library(ggplot2)
library(cowplot)
Attaching package: 'cowplot'
The following object is masked from 'package:ggpubr':
get_legend
Attaching package: 'gridExtra'
The following object is masked from 'package:dplyr':
combine
Attaching package: 'data.table'
The following objects are masked from 'package:dplyr':
between, first, last
# read in the excel sheet for the data
malit<- readxl::read_xlsx('Source Data.xlsx',sheet = "FigureS4")
malit <- as.data.table(malit)
# create subset of columns
df <- data.frame(reference=malit$reference, man_group=malit$man_group, man_name=malit$man_name,
ind_code=malit$ind_code, type=malit$type,
moderator.factor=malit$moderator.factor, co.variate.label=malit$co.variate.label,
dyr.1=malit$dyr.1, SEyr.1=as.numeric(malit$SDyr.1), n.O=malit$n)
# weighted mean of each covariate group, higher weight where more observations
df.wm <- df %>% group_by(ind_code,man_group,man_name,co.variate.label,type=malit$type) %>%
summarise(dyr.1 = signif(weighted.mean(dyr.1, n.O), digits=2),
reference = paste0(reference, collapse = ", "), n.S = n(), n.O = sum(n.O))
`summarise()` has grouped output by 'ind_code', 'man_group', 'man_name',
'co.variate.label'. You can override using the `.groups` argument.
df.wm$co.variate.label <- factor(as.factor(df.wm$co.variate.label), levels=c(
"rice crop",
"maize crop",
"wheat crop",
"high N rate",
"medium N rate",
"low N rate",
"high SOC",
"medium SOC",
"low SOC",
"high soil pH",
"neutral soil pH",
"low soil pH",
"high Clay",
"medium Clay",
"low Clay",
"high MAT",
"medium MAT",
"low MAT",
"high MAP",
"medium MAP",
"low MAP"))
# overall weighted mean of all covariate groups for each measure-impact
range <- df.wm %>% group_by(ind_code,man_group,man_name) %>%
summarise(min=signif(min(dyr.1), digits=2), max=signif(max(dyr.1), digits=2), dyr.1 = signif(weighted.mean(dyr.1, n.O), digits=2), min.perc.diff = signif((((min-dyr.1)/dyr.1)*100),digits=2), max.perc.diff = signif((((max-dyr.1)/dyr.1)*100),digits=2))
`summarise()` has grouped output by 'ind_code', 'man_group'. You can override
using the `.groups` argument.
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Crop rotation plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Crop rotation")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Crop rotation")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nrot <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Cover cropping plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Cover cropping")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Cover cropping")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.ncc <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Residue retention plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Residue retention")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Residue retention")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nres <- as.data.frame(cbind(grand_mean, t))
#================ combine data for each indicator =============================
t1 <- rbind(t.nres,t.ncc,t.nrot)
#==============================================================================
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Reduced tillage plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Reduced tillage")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Reduced tillage")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nrt <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and No tillage plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Zero tillage")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Zero tillage")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nnt <- as.data.frame(cbind(grand_mean, t))
#================ combine data for each indicator =============================
t2 <- rbind(t.nnt,t.nrt)
#==============================================================================
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Organic fertilizer plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Organic fertilizer")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Organic fertilizer")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nof <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Combined fertilizer plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Combined fertilizer")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Combined fertilizer")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.ncf <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Right fertilizer rate plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Fertilizer rate")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Fertilizer rate")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nrfr <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Right fertilizer timing plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Fertilizer timing")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Fertilizer timing")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nrft <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Right fertilizer placement plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Fertilizer placement")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Fertilizer placement")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nrfp <- as.data.frame(cbind(grand_mean, t))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# NUE and Enhanced efficiency plot
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
#subset of data man/ind pair and columns needed
s <- subset(df.wm, ind_code == "NUE" & man_group=="Enhanced efficiency")
t <-data.frame(ind=s$ind_code,man=s$man_name,type=as.factor(s$type),group=as.factor(s$co.variate.label),group_mean=as.numeric(s$dyr.1))
#extract overall mean man/ind pair
i<-which(range$ind_code == "NUE" & range$man_group=="Enhanced efficiency")
grand_mean <- rep(range$dyr.1[i],length(t$group))
t.nee <- as.data.frame(cbind(grand_mean, t))
#================ combine data for each indicator =============================
t3 <- rbind(t.nee,t.ncf,t.nof,t.nrfp,t.nrfr,t.nrft)
#==============================================================================
t1$man <- factor(as.factor(t1$man), levels=c(
"Residue retention",
"Cover cropping",
"Crop rotation"))
t2$man <- factor(as.factor(t2$man), levels=c(
"Zero tillage",
"Reduced tillage"))
t3$man <- factor(as.factor(t3$man), levels=c(
"Enhanced efficiency",
"Combined fertilizer",
"Organic fertilizer",
"Fertilizer placement",
"Fertilizer rate",
"Fertilizer timing"))
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
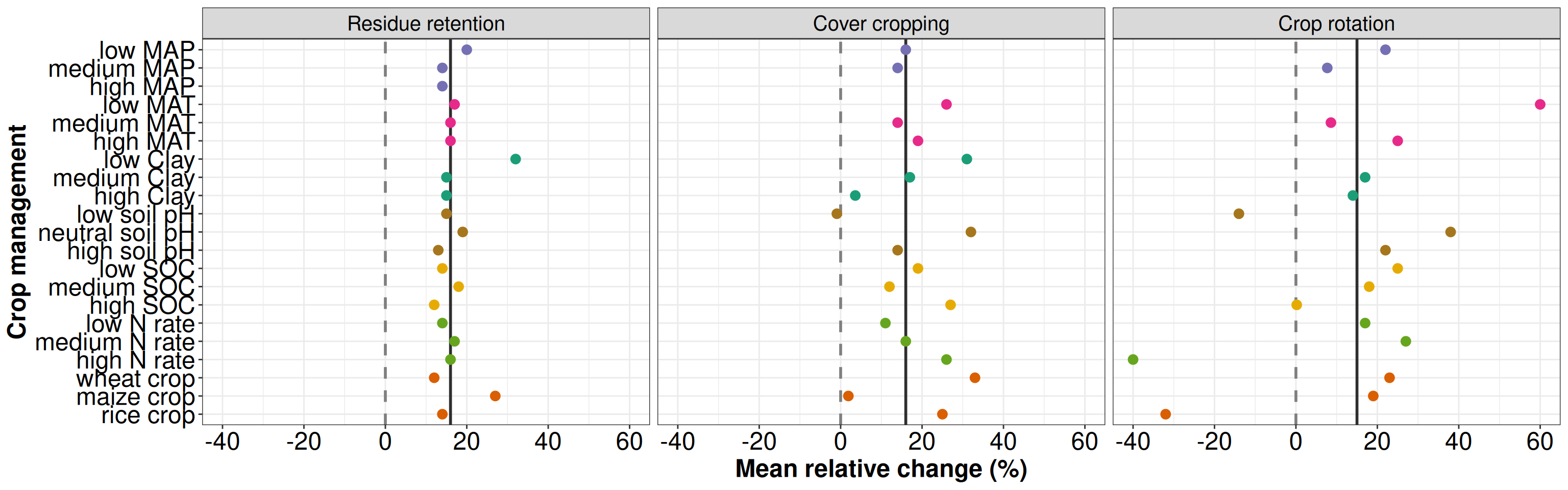
p1 <- ggplot(data = t1, aes(x = group, y = group_mean, color = type)) +
# add vertical line with grand mean
geom_hline(data = t1, aes(yintercept = grand_mean), linetype= 1, size=1, color="grey18", group= "man") +
# add vertical line for zero
geom_hline(yintercept=0, linetype = 2, size=1, color="grey50")+
# add points, flip coordinates and add own color scheme
geom_point(size=3) + coord_flip() +
scale_color_manual(values=c("#1b9e77", "#d95f02","#7570b3", "#e7298a", "#66a61e", "#e6ab02", "#a6761d"))+
facet_grid(cols = vars(man)) +
# add labels and general theme
xlab('Crop management') + ylab('Mean relative change (%)') + theme_bw()+
theme(axis.text.x = element_text(color="black", size = 18),
axis.text.y = element_text(color="black", size = 18),
axis.title = element_text(color="black", size = 18, face="bold"),
strip.text = element_text(color="black", size = 15),
plot.tag = element_text(color="black", size=30),
legend.position = "none")
Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
ℹ Please use `linewidth` instead.
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
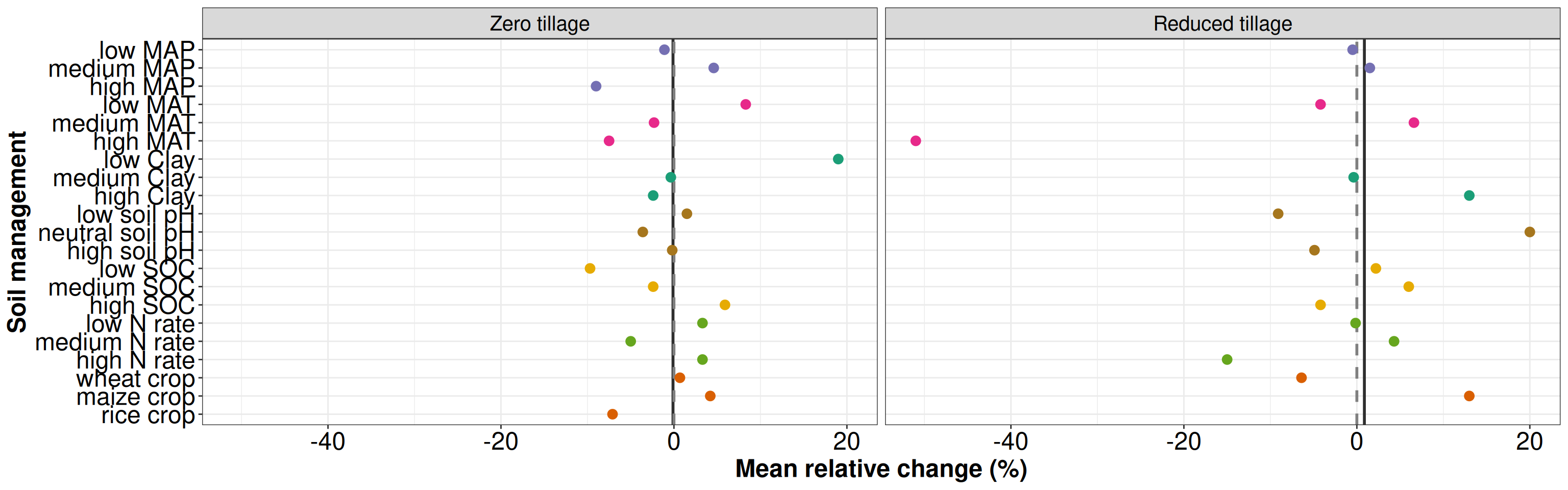
p2 <- ggplot(data = t2, aes(x = group, y = group_mean, color = type)) +
# add vertical line with grand mean
geom_hline(data = t2, aes(yintercept = grand_mean), linetype= 1, size=1, color="grey18", group= "man") +
# add vertical line for zero
geom_hline(yintercept=0, linetype = 2, size=1, color="grey50")+
# add points, flip coordinates and add own color scheme
geom_point(size=3) + coord_flip() +
scale_color_manual(values=c("#1b9e77", "#d95f02","#7570b3", "#e7298a", "#66a61e", "#e6ab02", "#a6761d"))+
facet_grid(cols = vars(man)) +
# add labels and general theme
xlab('Soil management') + ylab('Mean relative change (%)') + theme_bw()+
theme(axis.text.x = element_text(color="black", size = 18),
axis.text.y = element_text(color="black", size = 18),
axis.title = element_text(color="black", size = 18, face="bold"),
strip.text = element_text(color="black", size = 15),
plot.tag = element_text(color="black", size=30),
legend.position = "none")
p2
#++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
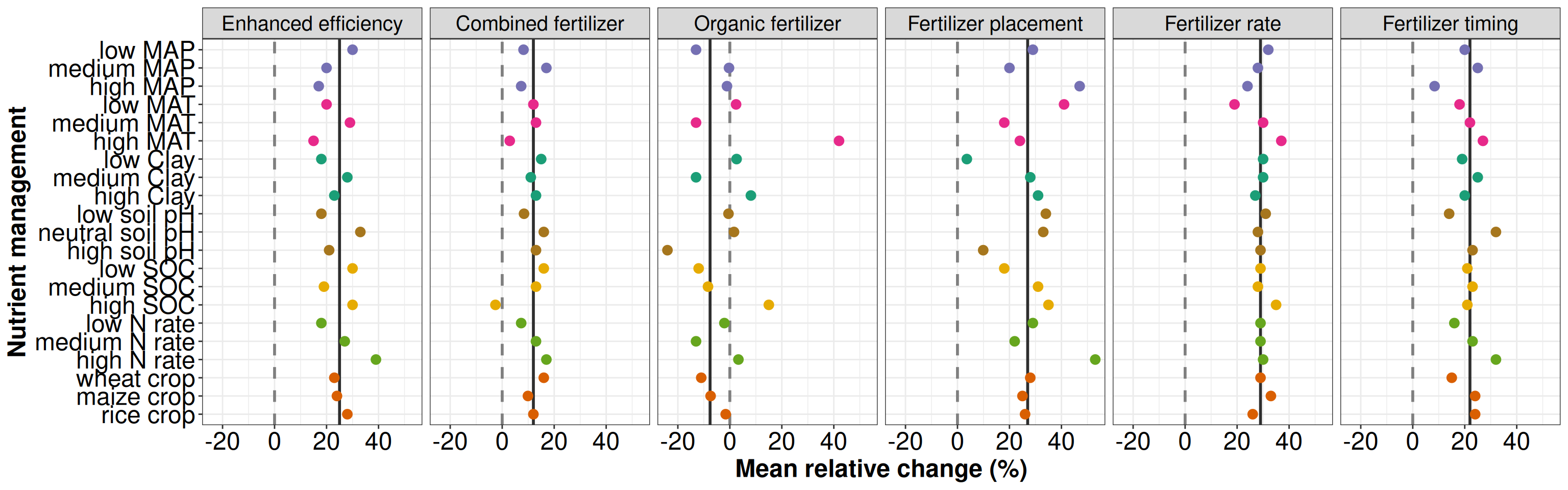
p3 <- ggplot(data = t3, aes(x = group, y = group_mean, color = type)) +
# add vertical line with grand mean
geom_hline(data = t3, aes(yintercept = grand_mean), linetype= 1, size=1, color="grey18", group= "man") +
# add vertical line for zero
geom_hline(yintercept=0, linetype = 2, size=1, color="grey50")+
# add points, flip coordinates and add own color scheme
geom_point(size=3) + coord_flip() +
scale_color_manual(values=c("#1b9e77", "#d95f02","#7570b3", "#e7298a", "#66a61e", "#e6ab02", "#a6761d"))+
facet_grid(cols = vars(man)) +
# add labels and general theme
xlab('Nutrient management') + ylab('Mean relative change (%)') + theme_bw() +
theme(axis.text.x = element_text(color="black", size = 18),
axis.text.y = element_text(color="black", size = 18),
axis.title = element_text(color="black", size = 18, face="bold"),
strip.text = element_text(color="black", size = 15),
plot.tag = element_text(color="black", size=30),
legend.position = "none")
p3